My is not distributed according to my wishes!
Let be a variable that we would like to model, for instance, Covid-19 cases.
Now, there’s a widely hold belief that my must be distributed normally, or, in some cases, following some other assumed distribution (maybe some long-tailed distribution).
However, this belief is not (strictly) true. What a linear model assumes is that the residuals are distributed normally, not the distribution. While this is often left implicit, it is helpful to spell out a model explicitly.
Here’s a simple example from Gelmann and Hill, 2007, p. 46.
Consider a regression with the predictor , which takes on the values 0, 1, and 2, equally distributed:
where referes to a discrete uniform distribution with minimum and maximum .
Now, let the true association of and amount to:
Having specified this, we have completely specified this data generating process.
For modelling purposes, let’s add some noise:
Simulate some data
Let’s draw a sample of .
n <- 1000
set.seed(42)
X <- sample(x = c(1,2,3), size = n, replace = T)
e <- rnorm(n, mean = 0, sd = 0.1)Compute and :
Y <- 0.2 + 0.5*X
Y_hat <- 0.02 + 0.5*X + eAnd put it all in a data frame:
X <- tibble(
X = X,
Y = Y,
Y_hat = Y_hat,
e = e,
)Plot the data
X %>%
ggplot(aes(x = X)) +
geom_histogram() +
scale_x_continuous(limits = c(0,3))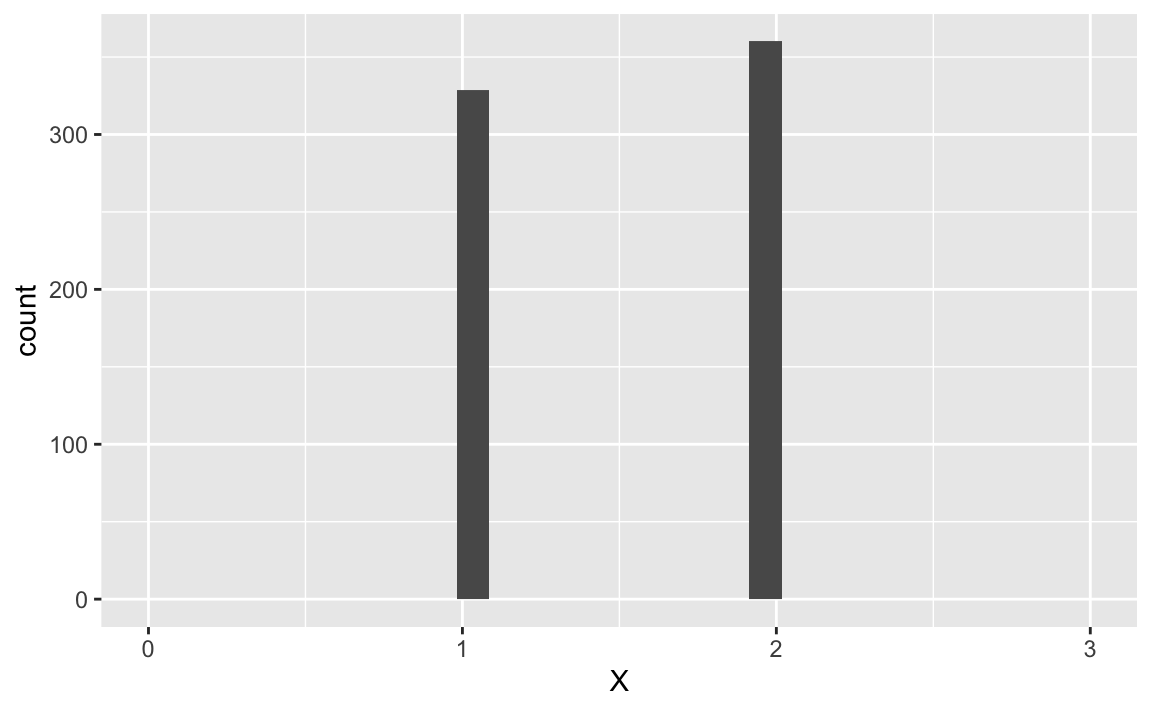
X %>%
ggplot(aes(x = Y)) +
geom_histogram() +
scale_x_continuous(limits = c(0,3))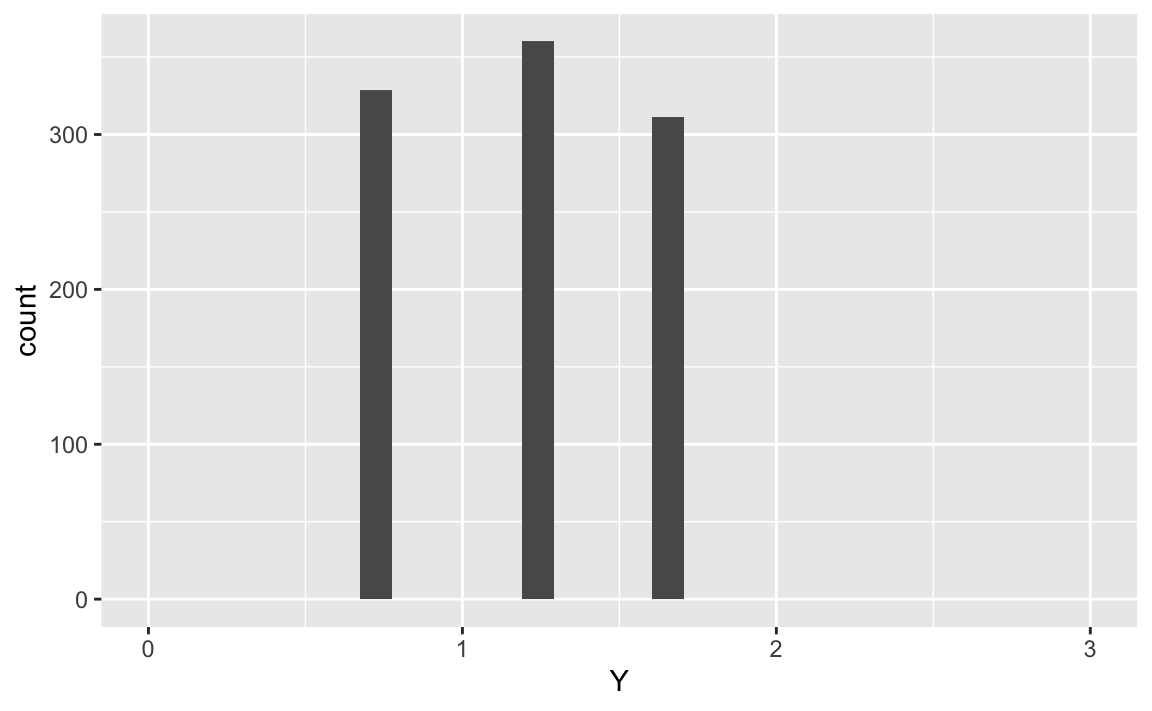
X %>%
ggplot(aes(x = Y_hat)) +
geom_histogram() +
scale_x_continuous(limits = c(0,3))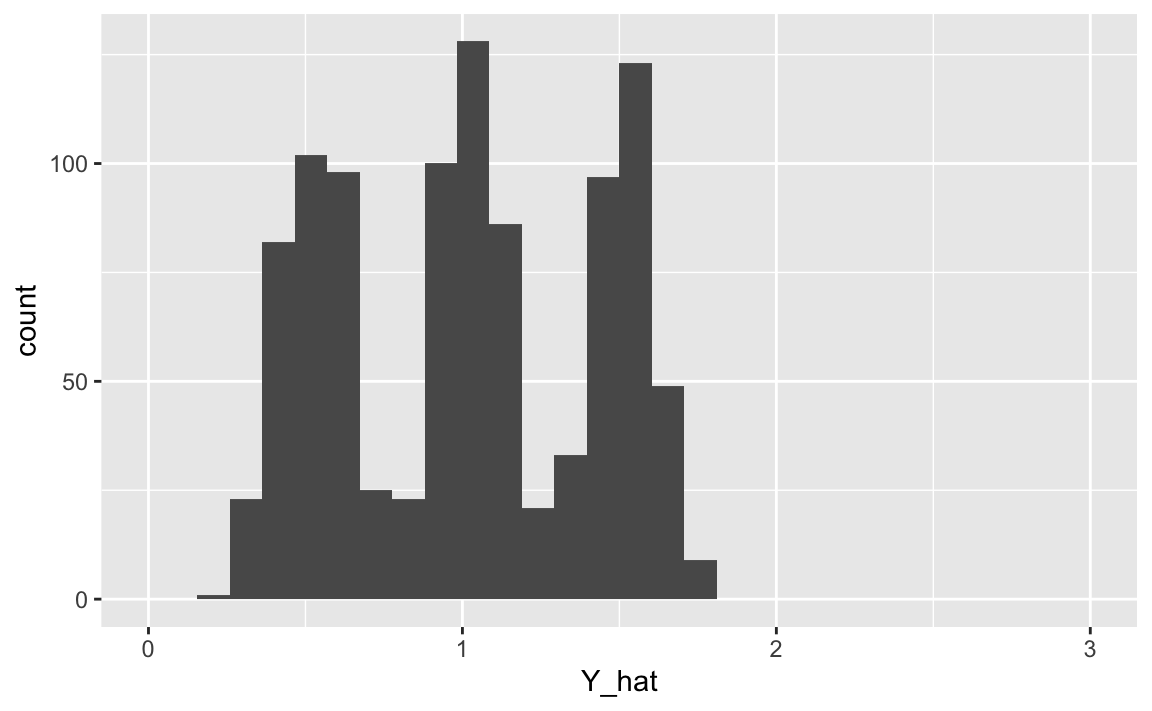
X %>%
ggplot(aes(x = e)) +
geom_histogram() +
scale_x_continuous(limits = c(-3,3))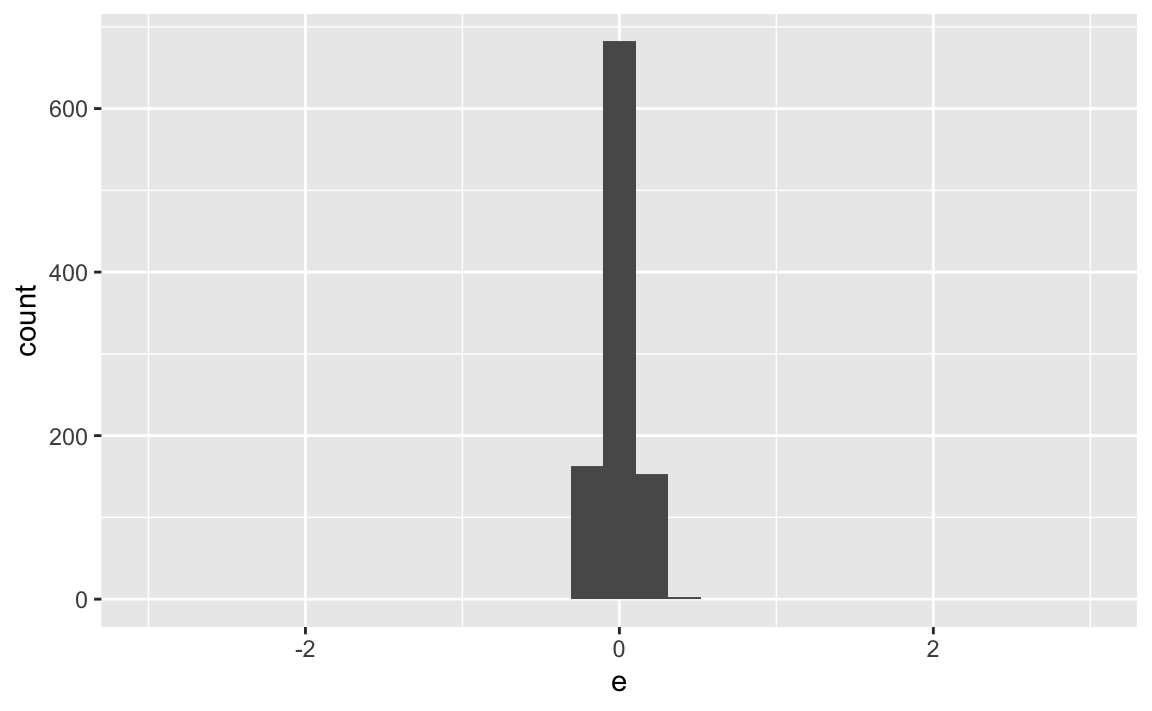
And interstingly, :
X %>%
ggplot(aes(x = e, group = X,
color = factor(X))) +
geom_density()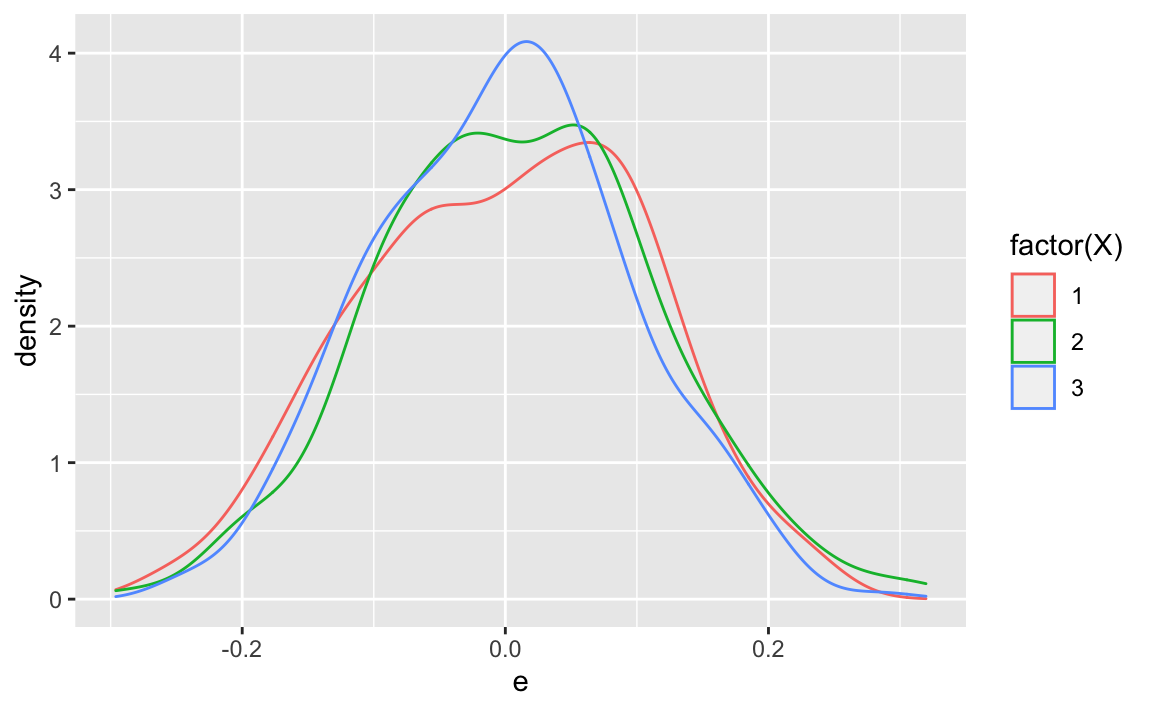
Compare that to , the unconditional (marginal) error distribution:
X %>%
ggplot(aes(x = e)) +
geom_density()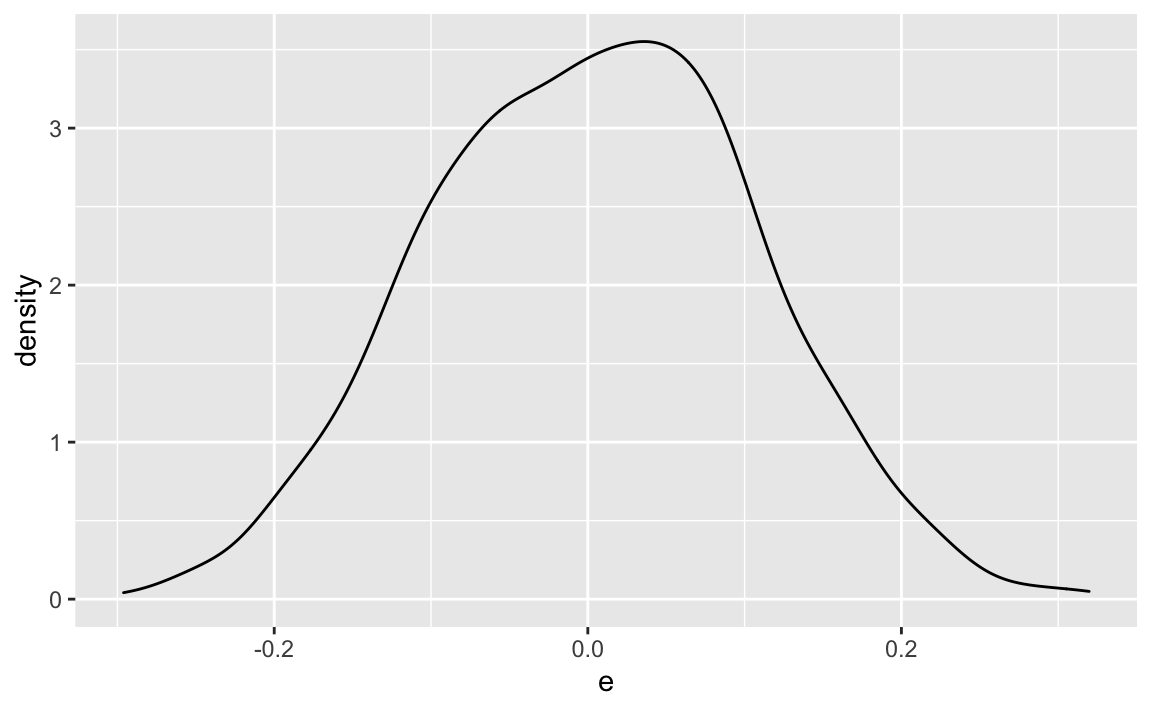
Quite similar, but: do we have thicker tails in compared to ? We have effectively mixed three distributions,so we might expect fatter tails.
e1071::kurtosis(X$X)
#> [1] -1.439171Recall that the kurtosis of the normal distribution is 3.
As a side note (same source as above):
It is also common practice to use an adjusted version of Pearson’s kurtosis, the excess kurtosis, which is the kurtosis minus 3, to provide the comparison to the normal distribution.
Some authors use “kurtosis” by itself to refer to the excess kurtosis. For clarity and generality, however, this article follows the non-excess convention and explicitly indicates where excess kurtosis is meant.
The function kurtosis provides the standard form of kurtosis, where normal distributed variables have a value of 3.
Compare that to the conditional kurtosis :
X %>%
group_by(X) %>%
summarise(kurtY = kurtosis(e))
#> # A tibble: 3 x 2
#> X kurtY
#> <dbl> <dbl>
#> 1 1 -0.615
#> 2 2 -0.0636
#> 3 3 -0.319There does not appear to be any difference in the kurtosis between and .
Conclusion 1
The distribution of is not of importance when it comes to the assumptions a linear models. However, the distribution of residuals (“errors”) is of some importance.